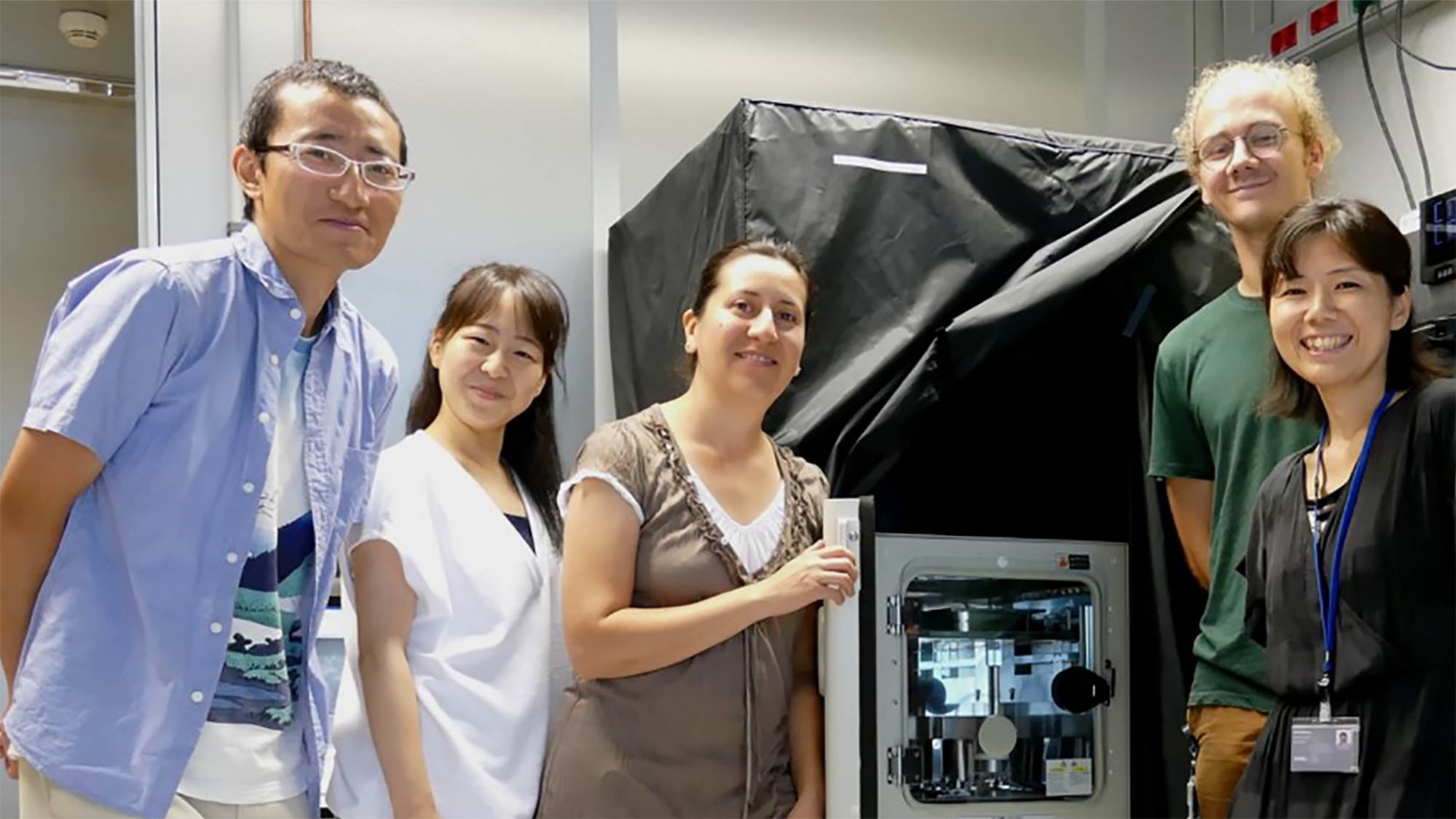
Miki Ebisuya started her lab at the European Molecular Biology Laboratory – Barcelona (EMBL Barcelona) a year ago, although it hasn’t been until this summer that her latest equipment and staff has arrived. “One of my postdocs (who is actually my husband) has been maintaining the lab in Japan during this year, finishing off things and shipping everything here. He finally joined me two months ago”, says Ebisuya.
Special equipment
The microscopes that have arrived at the 4th floor of the Barcelona Biomedical Research Park (PRBB) – where EMBL Barcelona is located – are not conventional. “We have a traditional confocal microscope, but also a custom-made microscope that we developed together with the Olympus company in Japan. That’s why we are waiting for this Japanese company to come and install it; the European branch of Olympus doesn’t sell it”, explains the head of the lab.
What makes this microscope special is that it detects bioluminescence, rather than fluorescence – which is the most common way of marking live cells.
Bioluminescence – what fireflies use to glow – doesn’t need any excitation. It’s an enzymatic reaction that catalyses the oxidation of the substrate luciferin, creating light. For the group, bioluminescence has some interesting advantages.
“We use a custom-made microscope that we developed together with the Olympus company in Japan, that detects bioluminescence, rather than fluorescence”
“One thing we are studying are oscillations (genes switching on and off in cycles), and these are seen as pulses of light (when a specific gene is expressed, the bioluminescence signal increases). Because the period of the cycles we are studying is short (they repeat every 2-5h), we can’t use fluorescent proteins, like GFP, because they have a half life of ~24h! So when the next pulse comes, we would still be seeing the light of the first one… And if we destabilize the fluorescent protein, the signal becomes too low to detect…,” explains the head of the group. Luckily, bioluminescence doesn’t produce background noise like fluorescence does, so the exposure time can be extended up to to 5-10min. This means that even if the signal is low, they can still detect it.
“Making” biology to understand the principles
The lab focuses on synthetic developmental biology. One half of the lab is trying to reconstitute developmental mechanisms as artificially as possible, while the other half tries to recapitulate developmental mechanisms in vitro, using stem cells. In both cases, the aim is the same: to understand the principles behind these developmental mechanisms using in vitro systems. They have been working in four processes:
1- Synthetic symmetry breaking
2- Synthetic pattern formation
3- Synthetic tissue deformation
4- In vitro oscillation
One half of the lab is trying to reconstitute developmental mechanisms as artificially as possible, while the other half tries to recapitulate developmental mechanisms in vitro, using stem cells.
1. Synthetic symmetry breaking
The first two processes were done while Ebisuya was still in Japan. For the first one, synthetic symmetry breaking, they used CHO cells (a common type of cultured hamster cells), which don’t have an endogenous intercellular symmetry breaking mechanism. The researchers introduced an artificial gene circuit based on Delta-Notch signalling (known to be involved in asymmetric cell division) and managed to get these cells to divide into two cell types expressing different markers. With this experiment the group has proved that the Notch system is not only necessary, but sufficient to produce this symmetry breaking. Recreating the phenotype artificially is the ultimate proof of the mechanism behind it. “And we also learned something new, since some components of the system didn’t behave in the expected way,” adds Ebisuya.
“Recreating a phenotype artificially is the ultimate proof of the mechanism behind it”
2. Synthetic pattern formation
Patterns are basic in development; for example our hand follows a pattern (finger, space, finger, space, finger, space, and so on until five). The second project of the lab was to recreate a synthetic cellular pattern. This time they used mammalian HEK293 cells where they introduced a gene circuit that represents the reaction-diffusion system, a mathematical model that explains how interaction (reaction) of multiple diffusible molecules can give rise to a spatial pattern.
With this new circuit, cells started to communicate with each other by sending signals which activate their neighbours and inhibit those cells that are a bit farther away. This creates a pattern of positive and negative areas (areas where a reporter gene is expressed or inhibited). Again, the reaction-diffusion system has been known for a while to be involved in this process, and computer simulations have been testing it for a long time – but “creating it from scratch, with real cells, genes and proteins, is a different thing,” says Ebisuya.
3. Synthetic tissue deformation
Both symmetry breaking and pattern formation are essential for development, but they are 1 or 2 dimensional mechanisms. Development occurs in 3D, so now the group is trying to manipulate the shape of tissues. They are using epithelial cell sheets on a gel with the aim of bending this sheet. As with the previous projects, what the group wants is to test the sufficiency of the known knowledge (in this case apical constriction) and to find out new mechanisms.
4. In vitro oscillation: the “Elephant time, Mouse time” problem
Another part of the lab is using stem cells to recapitulate natural developmental mechanisms, rather than reconstituting them from scratch. “It’s still in vitro” smiles the group leader, “just a bit less artificial”.
She remembers reading a Japanese book entitled Elephant time, Mouse time while at high school, that taught her that different species have different time cycles; larger animals live longer, have a slower heart beat, a longer gestation period… “Regretfully it didn’t tell me why, so the question stuck with me,” recalls the biologist.
Different species have different time cycles; larger animals tend to have longer time cycles but it’s not known why. Using stem cells, the lab is trying to recapitulate these mechanisms from different species in vitro, and compare them in the same conditions.
“I always thought this was too difficult a problem to study, because it’s complicated to compare what happens in different species when we have such different body environments; different cell numbers, different nutrition, etc. But now, thanks to induced pluripotent stem (iPS) cells and embryonic stem (ES) cells, we can recapitulate these mechanisms from different species in vitro, and then compare them in the same conditions in cell culture,” says Ebisuya.
So now her group is studying the segmentation clock, a type of oscillation that takes place during development. Interestingly, the oscillation tempo of the human segmentation clock is known to be slower than that of mice. The lab uses mouse and human pluripotent stem cells to recapitulate the segmentation clocks in vitro, studying the cause of the inter-species difference in biological time. “Our ultimate goal would be to modify something in human cells to get them to have the same time cycle than mouse, and otherwise. This would mean we understand the principles behind this difference”, says the Japanese researcher.
The Barcelona-Japan link
Ebisuya is quite settled in Barcelona. “I am still impressed by the beauty of the PRBB and the very interesting groups that are here,” she says. But that does not mean she is forgetting her origins.
“I am still impressed by the beauty of the PRBB and the very interesting groups that are here”
Miki Ebisuya
In an attempt to both keep in contact with her former colleagues in Japan, and give them the opportunity to meet her new neighbours, she has organised the “Japan-Barcelona Meeting on 3D Morphogenesis”, which will take place at the PRBB on October 21-22, 2019. Ebisuya encourages everyone at the PRBB, and the wider research community in Barcelona, to attend. “It’s free, but you need to register through the website!”, says the head of the group.

The young group leader further explains the motives behind the meeting: “Japan is a very isolated country, and because of our geography and the language barrier, it’s very difficult to communicate with researchers from other parts of the world… But there are many interesting researchers there – and I found many interesting researchers in Barcelona. So I wanted to bring the two groups together!”.
Ebisuya encourages everyone at the PRBB, and the wider research community in Barcelona, to attend the upcoming Japan-Barcelona Meeting on 3D Morphogenesis.
Building a community
Ebisuya is indeed interested in creating communities. At the PRBB, she has found good collaborators, such as Jordi Garcia Ojalvo, from Department of Experimental and Health Sciences, Pompeu Fabra University (DCEXS-UPF). “I knew about his work, but didn’t know he was in the same building! I actually thought he was in the US, because he collaborates a lot with American labs,” she admits. Garcia Ojalvo is her ideal collaborator, she says: he knows about oscillations and is quite familiar with synthetic biology. “He’s very busy, but because we are in the same building, I can come round his office and talk even if just for five minutes. This has facilitated our collaboration and we have even posted a preprint on bioRxiv already!” explains the scientist.
She is also part of a new monthly seminar series, the developmental biology club, with groups from EMBL, DCEXS-UPF, Centre for Genomic Regulation (CRG) and Hospital del Mar Medical Research Institute (IMIM). “It’s very good to know my neighbours!”, concludes Ebisuya.