I met Mar Albà in October, 2013 as I began my MSc in Bioinformatics at the Universitat Pompeu Fabra. She was the professor of a course called Principles of Genome Bioinformatics, where we learned some of the essential underlying concepts of bioinformatics, as well as getting some practical hands-on training.
When it came time to choose a project for my MSc thesis, I decided to join Mar’s lab after hearing about their research into finding out how de novo genes are born. Mar suggested that I read a really cool paper called “Proto-genes and de novo gene birth“, which proposed that hundreds of young de novo genes in baker’s yeast may have real adaptive potential. I was captivated by the mystery surrounding these de novo genes, and the potentially profound evolutionary consequences of their existence.
“I was captivated by the mystery surrounding de novo genes, and the potentially profound evolutionary consequences of their existence”
Will Blevins
For my thesis project, Mar proposed that we could do some experiments to follow up with the findings of the “Proto-genes…” paper: if we could assemble transcriptomes from scratch for a dozen closely-related species of yeast, we could dig even deeper into the question of where these young de novo genes came from.
More than just baking powder; yeast as a model system
I started to learn more about baker’s yeast, since I only knew a few things about it; I knew it was a single-celled eukaryote from the fungal kingdom, and that it was used to bake bread! However, I quickly learned that baker’s yeast AKA Saccharomyces cerevisiae could be used for much, much more than making bread.
I was impressed at how such a “simple” organism with only ~6000 genes had developed such ingenious and specific responses to changing environmental conditions. After reading more about S. cerevisiae, I still didn’t fully comprehend the true power of the experiments that one can design with a eukaryote that is so readily manipulated in the lab. This is where our collaboration with Lucas Carey’s Single Cell Behavior group, then at the Department of Experimental and Health Sciences, Pompeu Fabra University (DCEXS-UPF), began, since neither Mar nor myself had much experience designing our own experiments involving multiple species of yeast.
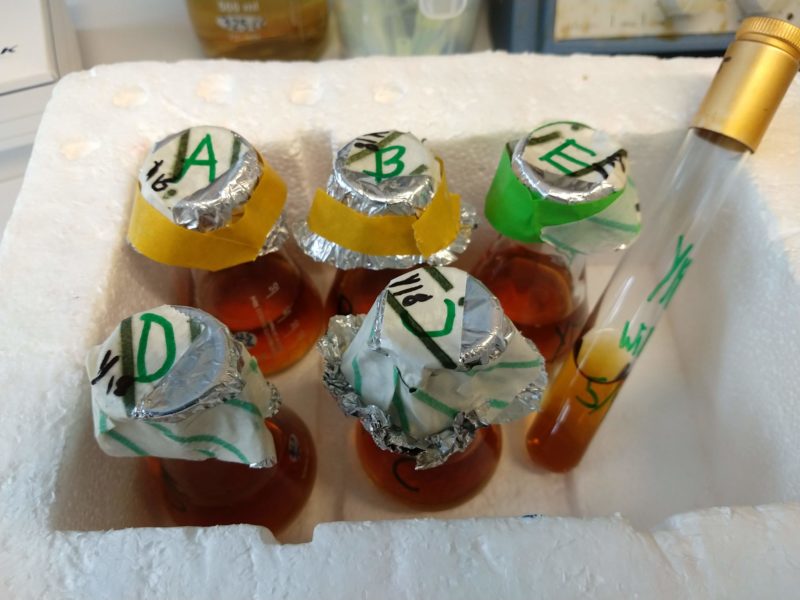

Together, we started to plan the experiments, trying to figure out the logistics and costs. Due to the amount of purified RNA we needed to be able to reconstruct the transcriptome for each species from scratch, we couldn’t run our experiments on 96-well plates (which would have made everything a LOT easier since you can automate many of the steps).
From dry to wet: what a computational biologist can learn in a lab
On my first day in the wet lab, I dropped “Yeast Stock Box #1”, sending a dozen cryo tubes flying, with a few cunning tubes hiding themselves under the lab bench. [To translate this for the computational folks out there, this would be like dropping an external HDD that contained the only copy of the raw data for the last 3 years of your projects.]

Fortunately, we were able to find all the tubes and put them back in the freezer before they could defrost, but I became *very* aware of how important it is to have physical backups… After all, there’s no CTRL+Z in the laboratory! Lorena Espinar was kind enough to guide me step-by-step through the protocols, going from frozen cells to purified RNA, which really helped me to better understand the raw sequencing data and potential sources of bias.
I experienced first-hand some of these common sources of bias that computational folks rarely encounter:
- media contamination (did you open the bottle without an open flame?)
- batch effects (did someone turn the shaker off overnight?)
- mislabeled samples (who entered this row in the spreadsheet?)
- missing data (where’s that post-it I used to write down the OD600 readings?)
Fortunately, after several months of tweaking and refining our experiments, we generated several pellets of purified RNA for each of our samples.
I experienced first-hand some sources of bias that as a computational researcher I had rarely encountered: media contamination, batch effects, mislabeled samples, missing data…
Back to computers
Coordinating the hand-off of our processed samples to the CRG-UPF sequencing facility felt like sending your child off to school for the first time – you know that they are in good hands, but you are still worried that something could go wrong.
After we got our raw sequencing data back, it was finally time to have a look at the last few months of hard work. We were happy to see that we had the depth and lower-limit of detection that we had aimed for, so we started to build our computational de novo gene-hunting pipeline. This analysis was the focus of my Bioinformatics Msc thesis project, and it was soon thereafter extended into a PhD thesis.

Expanding into a PhD
Spending 3-4 more years on this project meant that we could expand on some of our experiments, so we teamed up with Juana Díez’s Molecular Virology group, also at DCEXS-UPF, to find out which S. cerevisiae transcripts were being translated into peptides. Bernat Blasco had been working with a fairly new technique called ribosome profiling, and we were keen to collaborate.
“Attending the talks of visiting speakers, talking with other PhDs, and chatting with other PIs after conferences gave me some new perspectives on different sections of our analysis”

One of these interactions was with Xavier Messeguer from the Computer Sciences department at the UPC- he had previously developed a tool to recover syntenic regions between related species, and this collaboration helped to strengthen our overall methodology.
As the analysis progressed, we identified 213 transcripts which were only found in a few closely-related species and which had likely emerged de novo. About half of these taxonomically-restricted de novo transcripts were also being actively translated, meaning that these peptides had likely first appeared sometime over the last… ~20 million years! We also discovered that about half (105/213) of these de novo transcripts were found in an overlapping antisense orientation to other genes; this configuration was very uncommon for more conserved genes. You can read Mar’s behind-the-paper piece in Nature or this tweetorial to know more about this work!
Moving on in collaborative mode
There were many other ideas that we would have liked to follow through, such as some knockout experiments to test if these de novo transcripts contributed to the yeasts’ fitness, but unfortunately for various reasons, these ideas were never fully realized. As the date of my PhD defense grew ever closer, we wrote up a draft of the manuscript and submitted it as a preprint to bioRxiv.
It took over a year and several rounds of revisions, but in the end our article was finally published! I was very pleased, since this meant that the work from my PhD thesis would have a much broader impact, and it also meant that the people who made this work possible would get recognition too!
I think that these types of projects, which involve the collaboration of many different groups across different institutions, are crucial to advancing our understanding. I feel very fortunate that Mar is very adept at finding synergistic ways to join forces with other groups, and that in the Barcelona Biomedical Research Park (PRBB) there is a spirit of open collaboration, which helps us do better research and makes us all better researchers.
Blevins WR, Ruiz-Orera J, Messeguer X, Blasco-Moreno B, Villanueva-Cañas JL, Espinar L, Díez J, Carey LB, Albà MM. Uncovering de novo gene birth in yeast using deep transcriptomics. Nat Commun, 2021; 12(1): 604. PMID: 33504782 . DOI: 10.1038/s41467-021-20911-3.