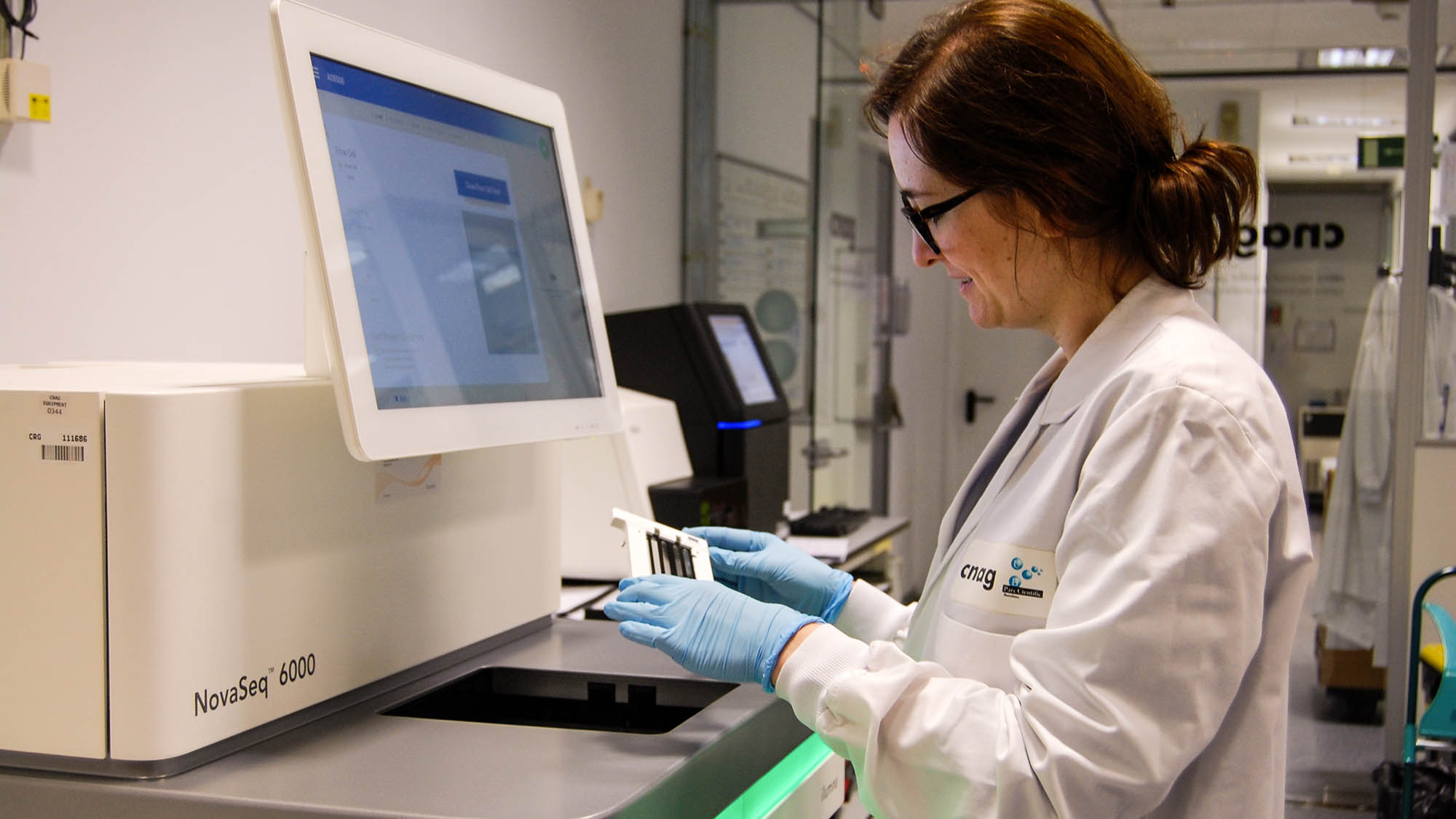
About 1,300 scientists from 70 research centres in 37 different countries have united to sequence and analyse the genomes from 2,658 donors with 38 types of tumors.
The result – 800 terabytes of information that represent the most comprehensive genomic map of 38 different types of tumours – has been published in 23 articles in the Nature journals.
This Pan-Cancer Atlas is the result of the analyses of data generated during more than a decade by The Cancer Genome Atlas (TCGA) and the International Cancer Genome Consortium (ICGC). Those previous projects had focused primarily on the 1% of the human genome that codes for proteins. Now, the researchers from the ‘Pan-Cancer Analysis of Whole Genomes (PCAWG) Project’ have looked into the remaining 99% of the genome to identify common patterns of mutations that could cause cancer – and therefore find new ways to prevent, diagnose and treat this disease.
In Spain, the Centro Nacional de Análisis Genómico (CNAG-CRG) has contributed, amongst other things, by sequencing primary tumours of chronic lymphatic leukaemia.
We speak to Ivo Gut, director of the CNAG-CRG.
The Pan-Cancer Genome Project has resulted in a comprehensive genomic map of 38 different types of tumours. Spain has focused in chronic lymphatic leukaemia.

What was the CNAG-CRG contribution to the project?
The Spanish contribution to the ICGC focused on chronic lymphatic leukaemia. Many hospitals were involved in procuring samples, coordinated from the Hospital Clínic. And several universities and research centres played an important part. At the CNAG-CRG, we sequenced roughly 100 samples of the Pan-Cancer project, which represent about 5%.
Apart from this, I think we made another important, though less visible contribution. The whole consortium was divided into 18 working groups. I led the last one to be created but, in my view, one of the most important: the Quality Control working group. Our aim was to ensure that all the sequences – which were coming from sequencing centres around the world – had sufficient quality. This was not so straightforward; we soon realised that everyone was defining quality in a different way! So we established 5 criteria, with specific thresholds, and determined those on all of the genomes. In the end that meant that we had to discard 10% of the 3000 genomes submitted, because they didn’t reach the threshold. Evidently, if you have lousy quality data, all the interpretation that comes out of it is not reliable…
What have been the challenges and highlights of this collaboration?
The sheer amount of data generated – each genome we analyse is 100.000.000.000 bases sequenced! It takes months to just copy all of the data from the over 2,500 cancer genomes. Just the effort of bringing all the data to one place is already a huge technical challenge. But in the end, what’s important is that we have proved that we can do it, and this initial work opens up endless possibilities for further studies.
For each patient, we obtained cancer tissue and healthy tissue, and compared their genomic sequences, to look for differences. But the beauty of this project is that then you can compare whatever you want! You can compare tens of genomes of the same type of tumour. Or you can compare different types of tumours with each other – to see if there are any mutations that are common to all types.
Just the effort of bringing all the data to one place is already a huge technical challenge. But this sheer amount of data oppens up endless possibilities for further studies.
What’s next in store?
I think in the near future we are going to see genomic information being commonly used in clinical decisions. The cost of sequencing is not so big – I think that in less than 5 years most people with cancer might have their genome sequenced. The findings of the Pan-cancer study, and everything that we will learn from the data, are key for the development of personalized medicine.
“In the near future we are going to see genomic information being commonly used in clinical decisions”
There are new studies on the same line. I am currently participating in the “1+Million Genomes”, a European member states initiative, where 21 countries across Europe are trying to establish a system by which they can share genomic and clinical information with each other. Each country holds its own information, but allows qualified individuals to search across the dataset. The idea is to have 1 million genomes available by 2022, and eventually integrate this into healthcare. This will be tremendously useful, all this data will help inform diagnosis and treatment, in particular in areas such as rare diseases and cancer.