The Centre for Genomic Regulation (CRG) laboratory led by Eva Maria Novoa has recently published a new methodology that allows, for the first time, to quantify the levels of transfer RNA (tRNA) and its modifications quickly and inexpensively. This opens the door to a possible revolutionary diagnostic test for cancer that could, in less than 3 hours and for less than €50, indicate who has cancer and what type – all from a simple blood sample.
Cancer detection: DNA vs RNA
Until now, apart from imaging or other clinical tests, cancer could be detected from DNA (by sequencing existing mutations) or from circulating mRNA in the blood, released through the rupture of tumour cells. DNA sequencing has been done routinely for decades, with Illumina being the most widely used technology. But “it can cost thousands of euros per sample,” explains Novoa.
RNA detection, on the other hand, can be more informative and cost-effective. RNAs from tumour cells are also released into the blood, and tRNAs are, in fact, the most abundant RNA molecules. Moreover, over the years, sufficient evidence has accumulated that RNA modifications are altered in many diseases, in particular in different types of cancer. Therefore, knowing which RNAs are in the blood, and with which modifications, could indicate the presence of tumour cells, as well as what type of tissue they come from. This is the idea behind the CRG group’s research.
tRNAs are the most abundant RNA molecules and their modifications are altered in many diseases, in particular in different types of cancer, making them very good cancer biomarkers.
RNA modifications, the crux of the matter
Oxford Nanopore Technologies, an alternative technology to Illumina, began in 2017 to allow sequencing not just of DNA, but of native RNA. “Until then, to know the amount of RNA with Illumina, you had to do reverse transcription (going from RNA to complementary DNA), then amplify the cDNA and sequence it. But a serious problem is that reverse transcription ‘erases’ the RNA modifications… so we could only know the abundance of RNA, but not the modifications. A lot of information is lost, important information,” says Novoa.
Illumina does allow to study RNA modifications, using specific antibodies against a particular modification each time, but this is indirect and predetermined, and without molecule resolution. And with RNAs that have many modifications – such as ribosomal (rRNA) or transfer RNA (tRNA), which can have about 30 modifications per molecule on average – it is almost impossible to convert them into DNA for sequencing by Illumina.
The advent of Nanopore technology made it possible to sequence not only DNA but also RNAs with their modifications, taking a more detailed ‘snapshot’ of the genetic information in the cell.
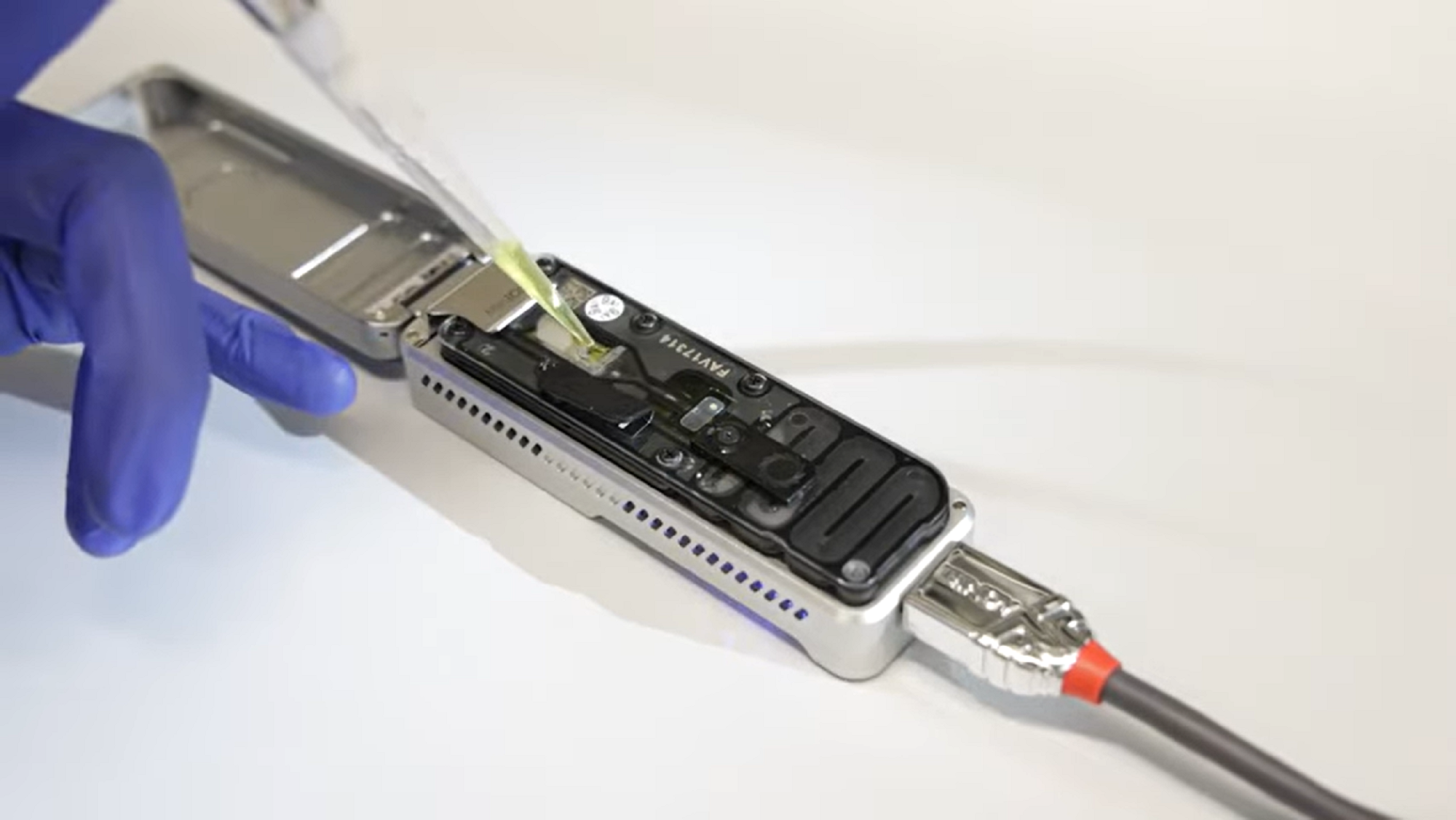
Nanopore, on the other hand, makes it possible to take a ‘picture’ of the RNAs in the sample as they are, with their modifications – and all of them at the same time. The technology consists of a small, hand-held device, about 10cm long, containing membranes with nanopores or channels through which RNA or DNA molecules pass. As they pass through the pores, they alter the electrical current passing through them, and the device reads this change in current and deduces the sequence and modifications. Some preparation is still needed (“you have to add an adaptor to the RNAs to make them pass through the nanopores at a constant, slower rate, which allows them to be read,” says the researcher), but it is fairly straightforward, and the amount of information obtained is very large.
Commercial Nanopore kits allow mRNA sequencing, and have long been used, because mRNA abundance can be used as a biomarker for some cancers. But mRNAs are not very abundant and many of them need to be sequenced to find differences between cancer and non-cancer samples. Non-coding RNAs (such as the tRNAs Novoa’s group works with), on the other hand, are much more abundant, and therefore much more informative and better biomarkers, with more diagnostic capacity.
The group led by Eva M. Novoa has succeeded in using Nanopore to sequence transfer RNAs, which are the most informative but very small and difficult to detect.
So, since the first Nanopore kits emerged, Novoa’s lab and others around the world have been making adaptations of this technology to allow sequencing of other types of RNA beyond mRNAs. One challenge with tRNAs, however, is that they are very short molecules – and therefore were lost and Nanopore did not manage to sequence them. “What we have been able to do now is to sequence these tRNAs well. We have shown that we can capture many molecules, detecting the correct abundances and their differential modifications,” says the group leader. This has been achieved thanks to modifications in the preparation of the libraries, and also in the computer analysis, in which algorithms and artificial intelligence are used to analyse the sequences and make predictions.
From proof-of-concept to clinical diagnosis
However, before this tRNA sequencing can be used for clinical diagnosis, it is necessary to better understand which specific modifications are linked to which type of cancer. “We know some of them, but they have not been analysed comprehensively, precisely because until now you couldn’t analyse these modifications in an ‘untargeted’ way; everyone looked at the 4-5 modifications that were known and were easier to look at,” says Novoa. But now they can take this ‘snapshot’, and this will be the first step of the group; to study and classify the modifications linked to each cancer.
To do this, the group is now looking for patient samples – breast and colon cancer to start with – both in biobanks and in collaboration with doctors in hospitals. “We need a variety of samples, from different types of cancer, and well stratified, to get the maximum information,” says the researcher.
Afterwards, the plasma sequencing technique will have to be optimised, as so far the studies have only been done with cell lines.
Finally, they will have to train the computational model, so that it can make predictions as the RNA is sequenced. “How far these predictions will go, we don’t know. It might just be able to tell whether the sample has cancer or not – although I think it will almost certainly be able to detect what tissue it comes from“, says the biologist. “Maybe it could tell if it’s metastatic cancer or not. Or if it’s resistant to a certain treatment. Or what stage it is in… it will all depend on the amount of information we have to train the model”.
The ultimate goal would be to use this technology for screening and early diagnosis, because the earlier the cancer is detected, the greater the probability of survival and the lower the cost to the health system.
The goal would be to use this technology for screening and early diagnosis of many types of cancer at the same time.
In addition to being a small and very portable device – which would facilitate its use in places without large infrastructures, as all it needs is a good computer and electricity – RNA sequencing with Nanopore technology would allow screening for many cancers at the same time. “When we do a mammogram, for example, apart from its cost, we are only detecting breast cancer. But by testing a blood sample with Nanopore, we think we will be able to see signs of any cancer. Afterwards, obviously, other tests should be done to validate the results, but it would be a great first step,” concludes Novoa.